Research Articles
Decoding Metabolism: A Comparative Guide to Flux Distribution Algorithms for Systems Biology
This article provides a comprehensive overview for systems biology researchers and metabolic engineers comparing the flux distributions predicted by different computational algorithms.
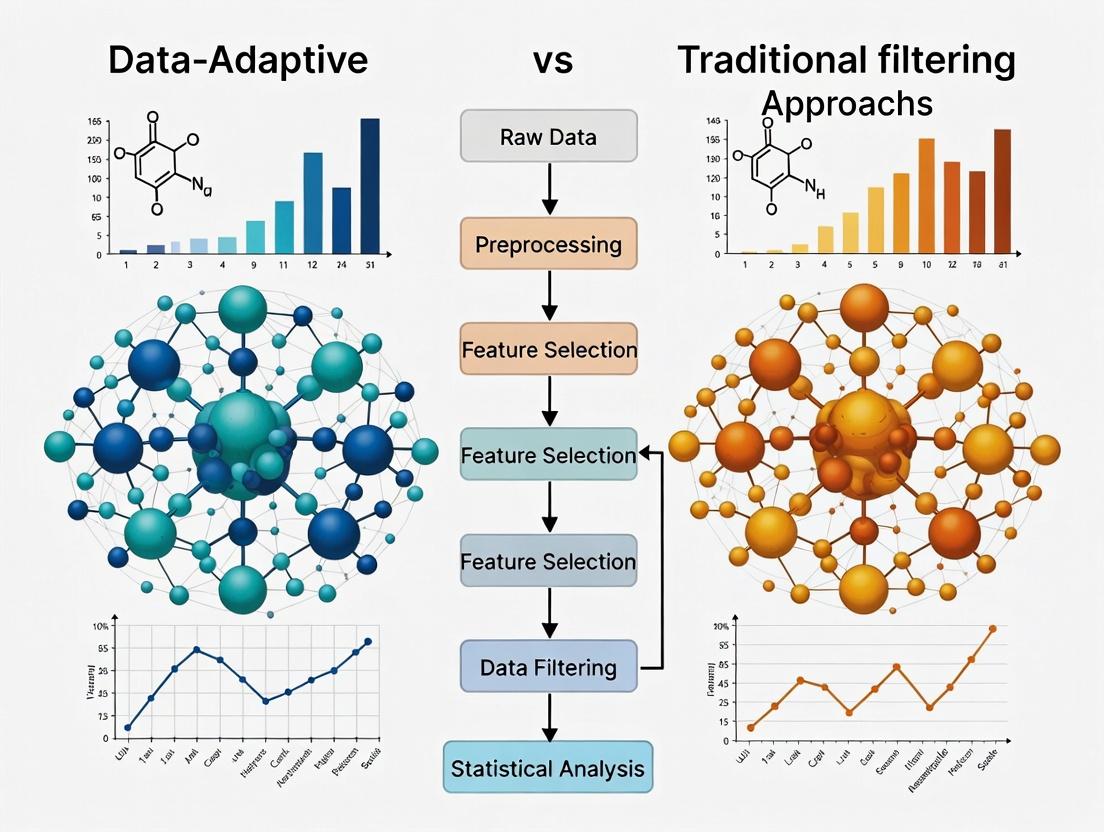
Filtering Revolution: Data-Adaptive vs Traditional Approaches in Modern Metabolomics Research
This article provides a comprehensive comparison of data-adaptive filtering versus traditional statistical methods in metabolomics data analysis.
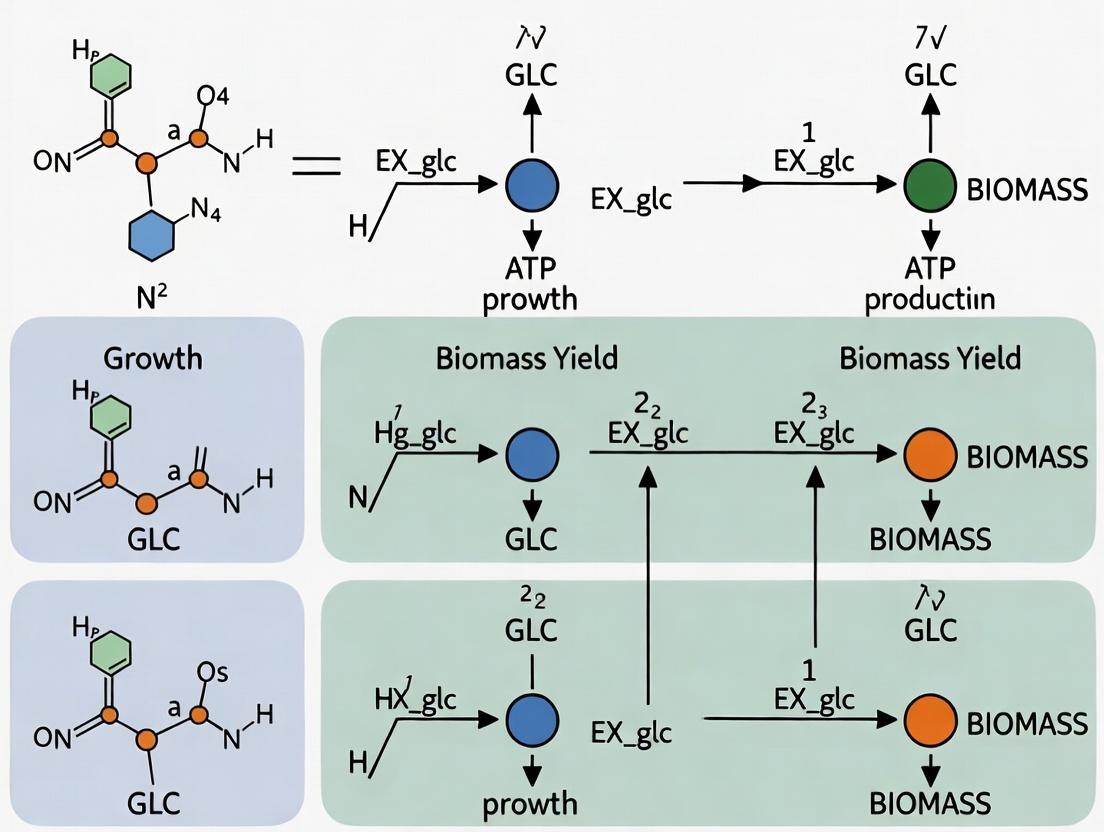
Unlocking Metabolic Predictions: A Comparative Guide to FBA Objective Functions for Underdetermined Systems
Flux Balance Analysis (FBA) is a cornerstone of constraint-based metabolic modeling, but its predictions hinge critically on the chosen objective function, especially for underdetermined systems with infinite flux solutions.
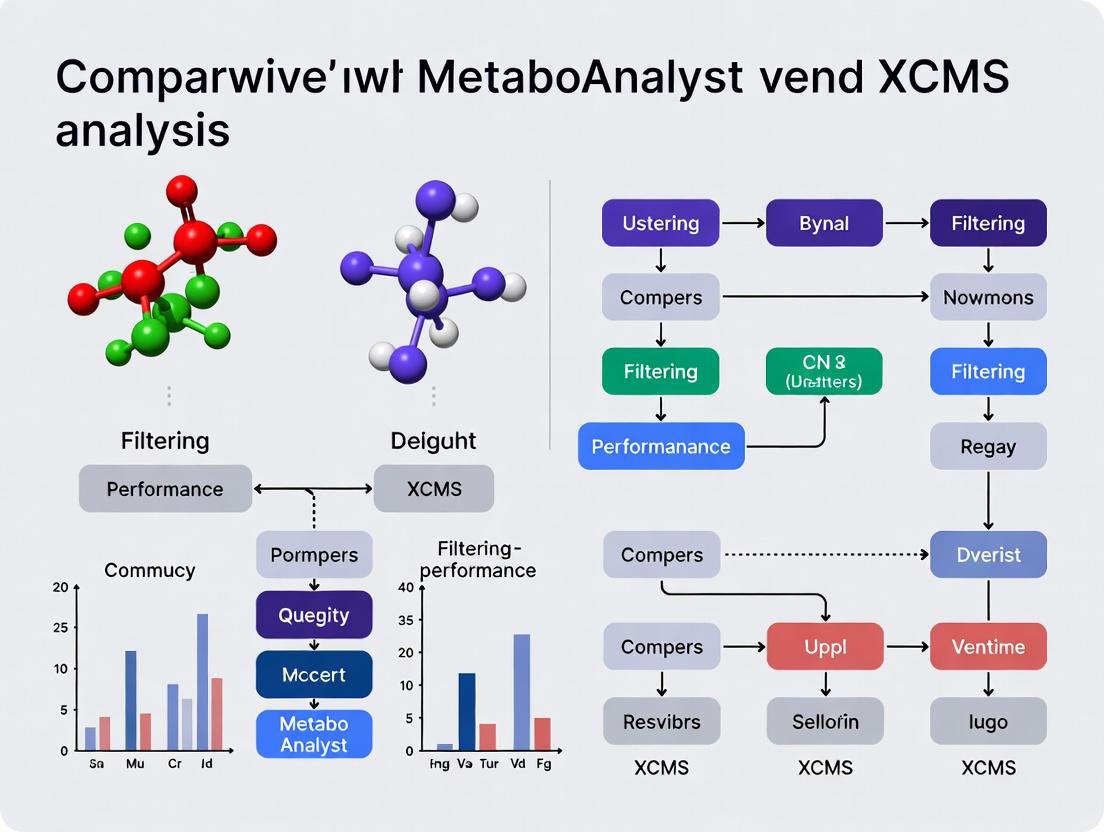
XCMS vs MetaboAnalyst: A 2024 Performance Guide to Peak Filtering for Precision Metabolomics
This comprehensive guide provides researchers, scientists, and drug development professionals with an up-to-date comparative analysis of peak filtering in XCMS and MetaboAnalyst.
Navigating the Noise: Overcoming Uninformative Features in Untargeted Metabolomics for Robust Biomarker Discovery
Untargeted metabolomics generates complex datasets rich with biological potential but plagued by uninformative features—chemical noise from contaminants, artifacts, and irrelevant biological variation.
Mastering Biomolecular Reconstruction: A Comprehensive Guide to CarveMe for Top-Down Metabolic Models
This tutorial provides a complete, step-by-step guide for researchers, scientists, and drug development professionals to master CarveMe for reconstructing genome-scale metabolic models (GEMs) from annotated genomes using the top-down approach.
A Guide to CarveMe Draft Model Reconstruction and Gap-Filling: From Theory to Practice for Drug Discovery
This comprehensive guide details the CarveMe reconstruction pipeline for generating genome-scale metabolic models (GSMMs), with a specific focus on automated draft model reconstruction and essential gap-filling strategies.
CRISPR/Cas9 Metabolic Engineering: A Comprehensive Guide for Researchers on Precision Genome Editing
This article provides researchers, scientists, and drug development professionals with a detailed framework for applying CRISPR/Cas9 to metabolic engineering.
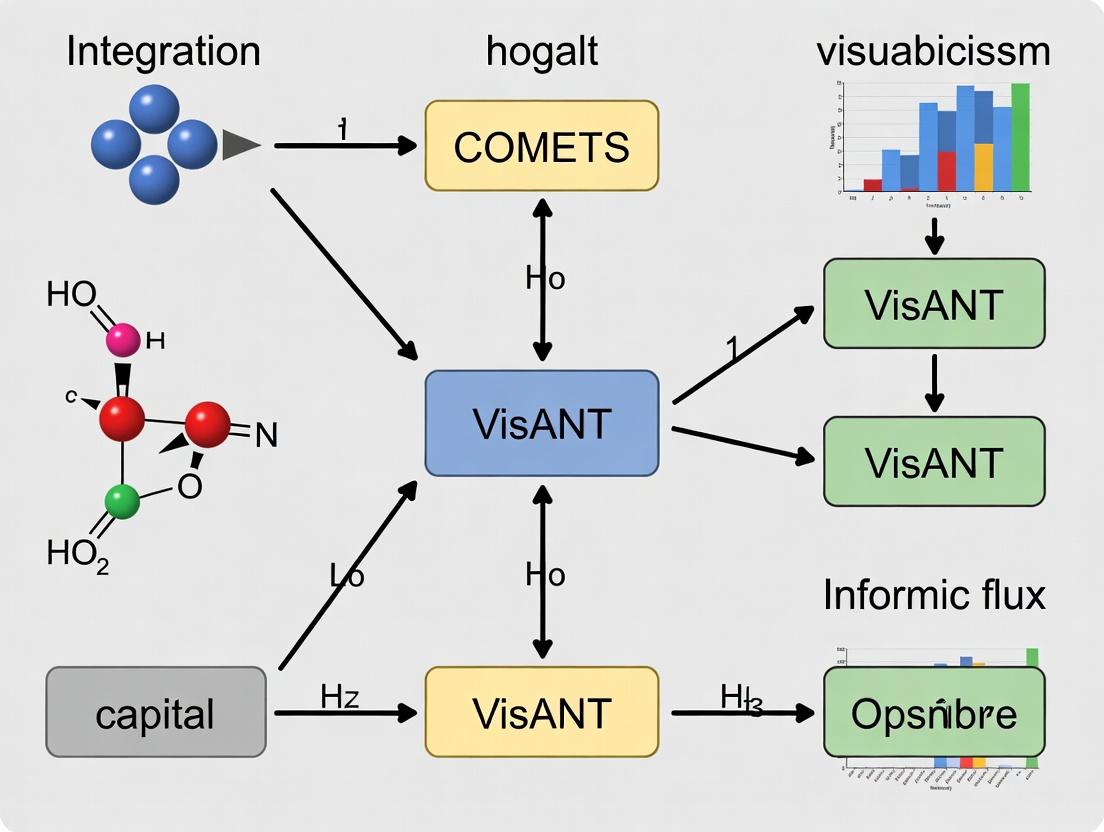
Dynamic Metabolic Visualization: Integrating COMETS and VisANT for Advanced Network Analysis
This article provides a comprehensive guide to integrating the COnstraint-Based Metabolic and Expression/regulation Task Space (COMETS) simulation platform with the Visual Analysis of Networks (VisANT) visualization tool for dynamic flux...
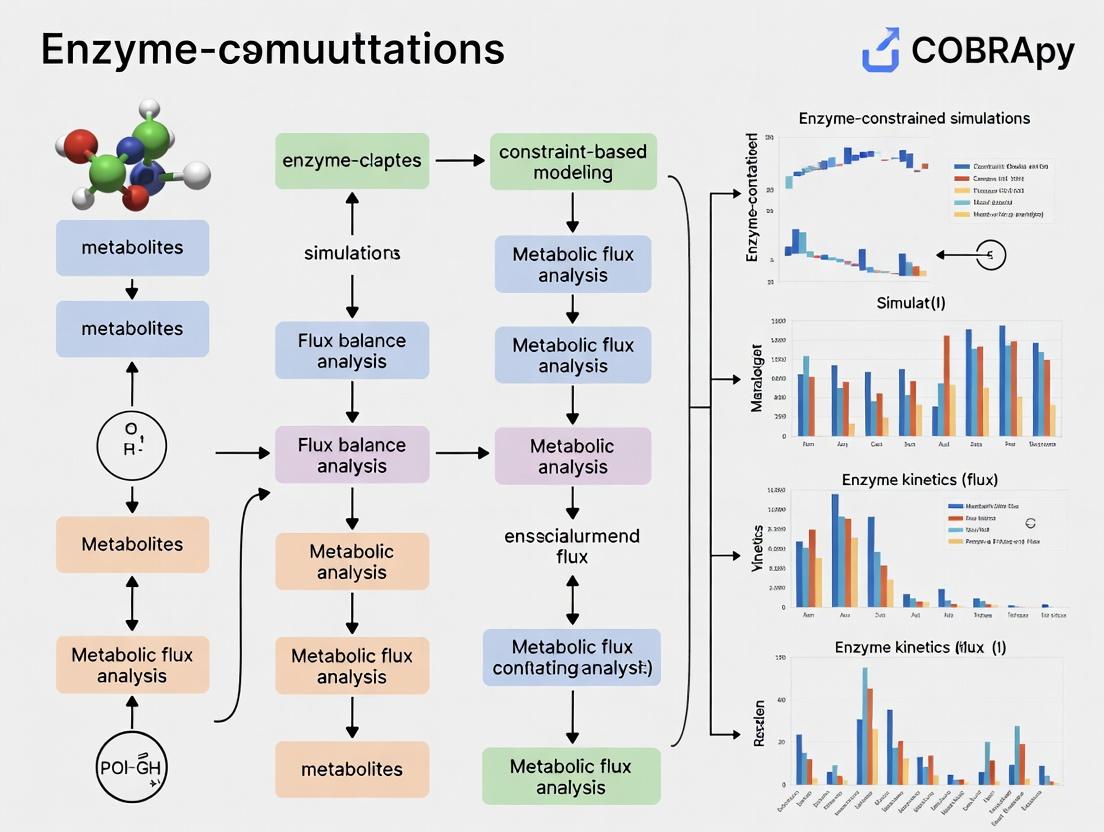
COBRApy for Enzyme-Constrained Modeling: A Complete Guide to ecFBA Simulations in Python
This comprehensive guide details the use of COBRApy to implement and apply Enzyme-Constrained Flux Balance Analysis (ecFBA) for metabolic modeling.